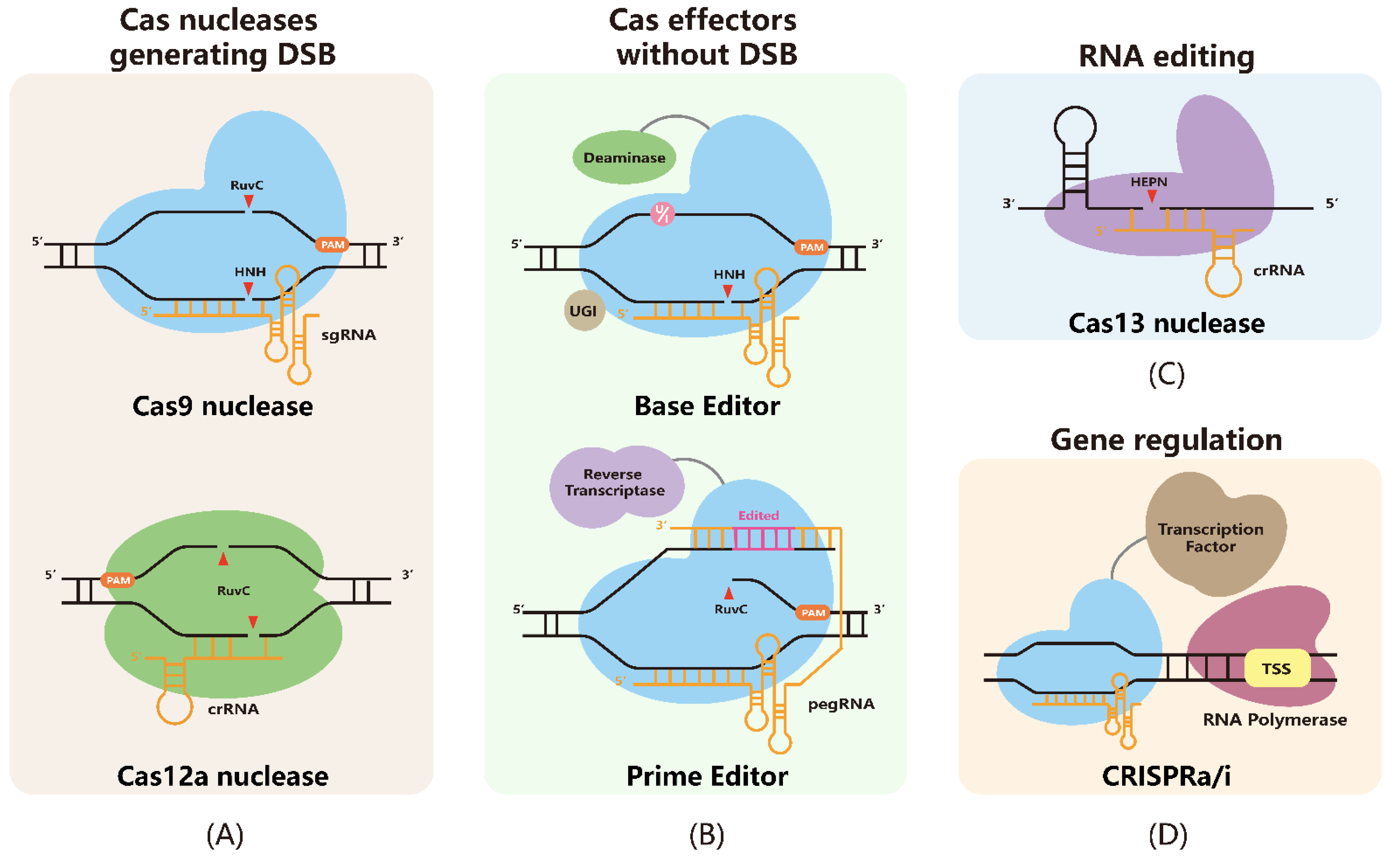
The application of CRISPR/Cas technologies to Brassica crops: current progress and future perspectives | SpringerLink

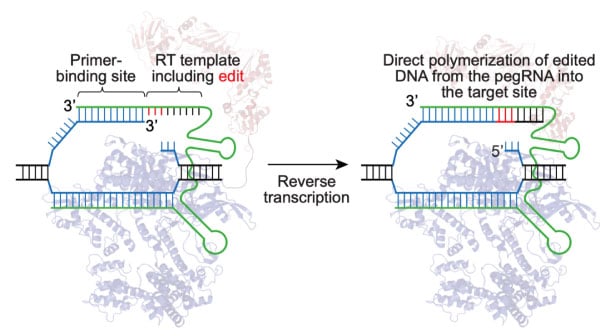
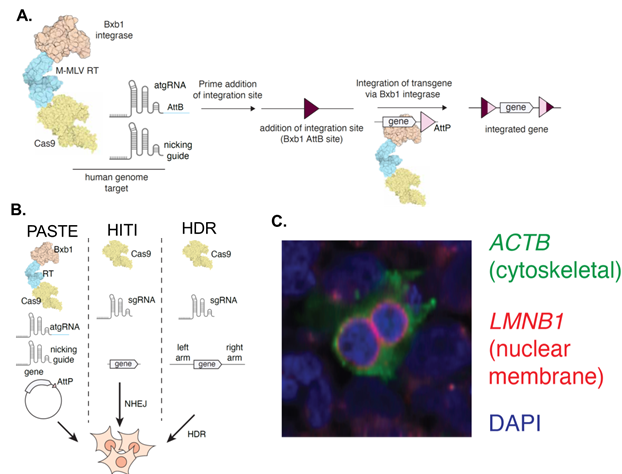
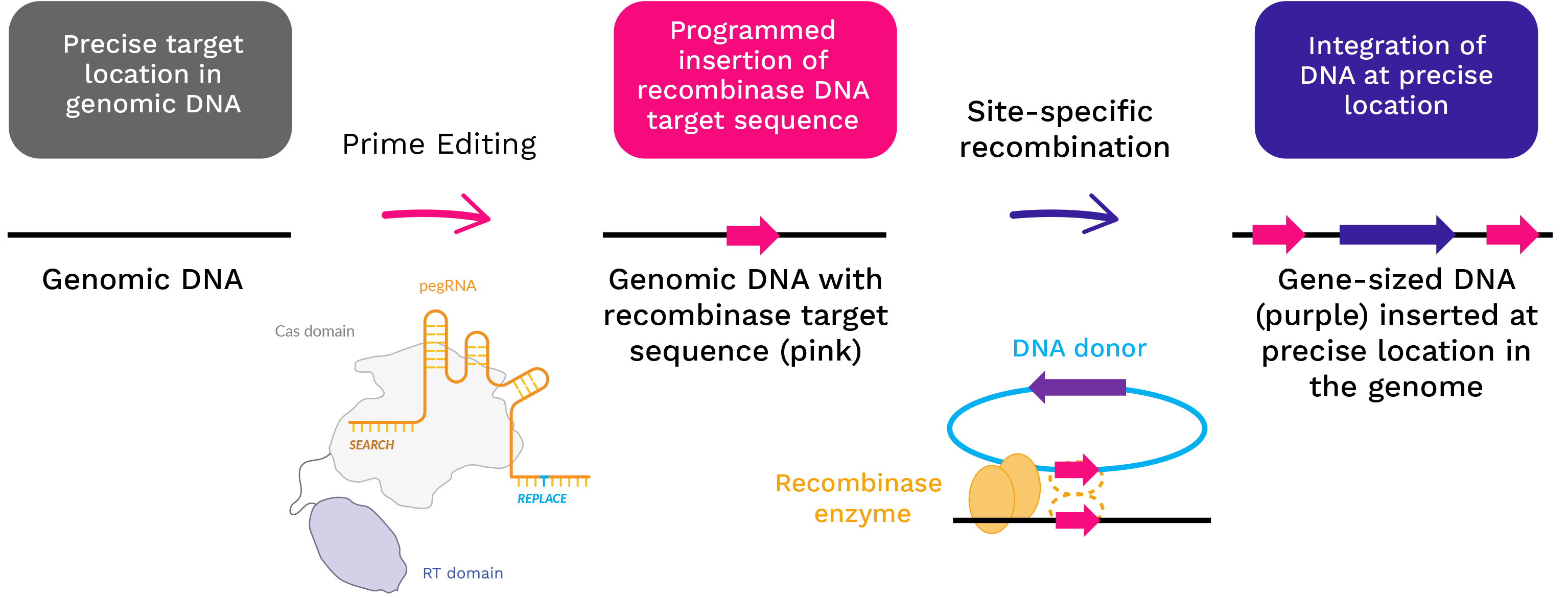
Programmable deletion, replacement, integration and inversion of large DNA sequences with twin prime editing | Nature Biotechnology

Programmable deletion, replacement, integration and inversion of large DNA sequences with twin prime editing | Nature Biotechnology

David R. Liu on Twitter: "TwinPE uses a prime editor protein and two pegRNAs to program the synthesis of complementary DNA flaps on opposing strands of genomic DNA, resulting in high-efficiency precise

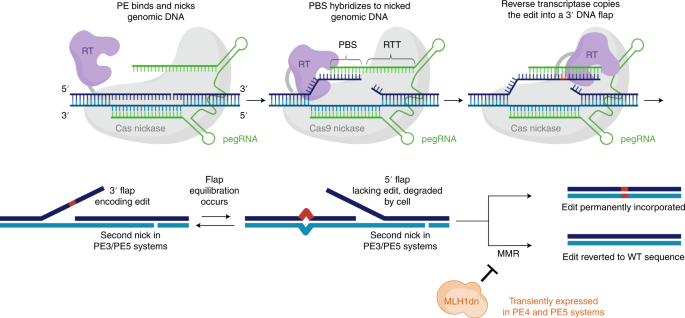
Cells | Free Full-Text | Future Perspectives of Prime Editing for the Treatment of Inherited Retinal Diseases

Prime editing techniques use two nicks. Prime editing (PE) uses a PE... | Download Scientific Diagram

Programmable deletion, replacement, integration and inversion of large DNA sequences with twin prime editing - ryr1.org

CRISPR Experts Introduce Twin Prime Editing- Crop Biotech Update (December 15, 2021) | Crop Biotech Update - ISAAA.org
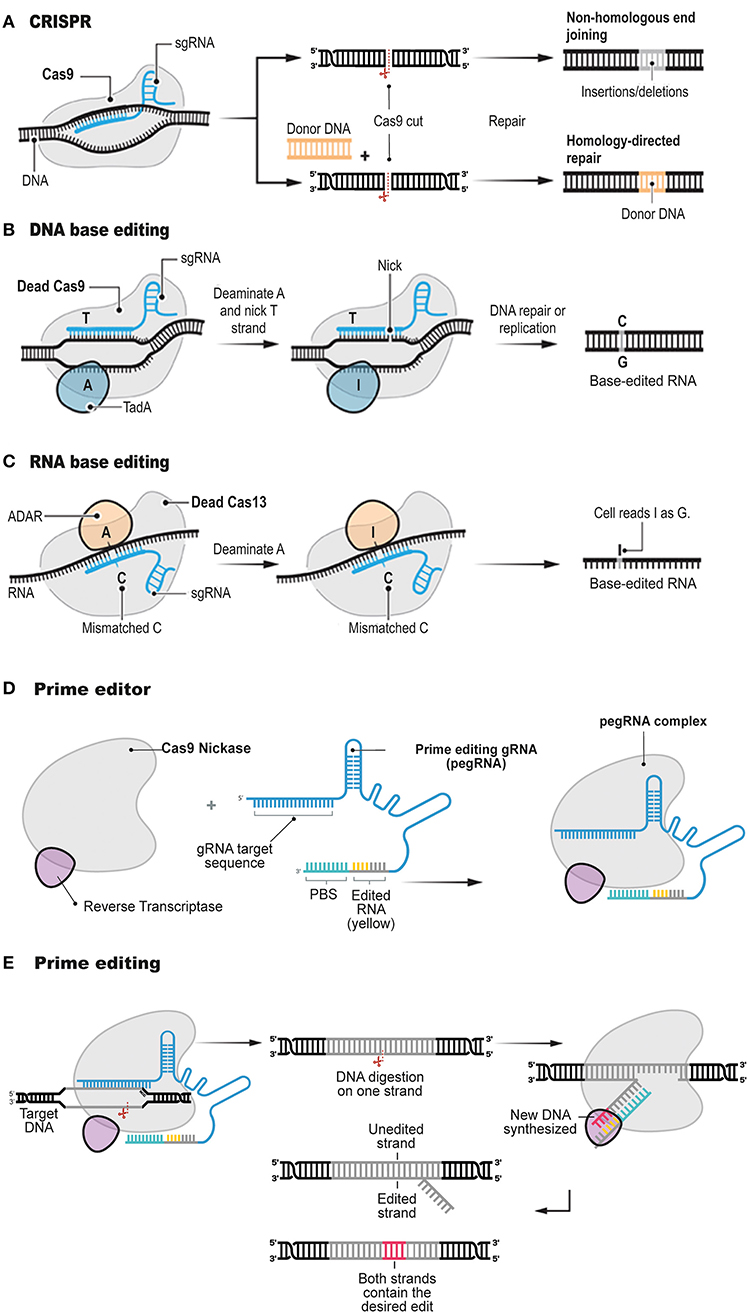
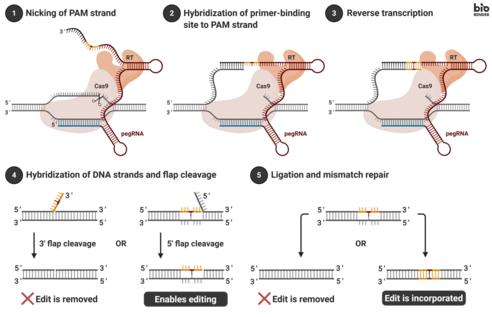
Frontiers | Prime Editing: Genome Editing for Rare Genetic Diseases Without Double-Strand Breaks or Donor DNA

David R. Liu on Twitter: "TwinPE uses a prime editor protein and two pegRNAs to program the synthesis of complementary DNA flaps on opposing strands of genomic DNA, resulting in high-efficiency precise

Programmable large DNA deletion, replacement, integration, and inversion with twin prime editing and site-specific recombinases | bioRxiv